Foundation cell segmentation models performance on live microscopy and spatial-omics data

Foundation cell segmentation models performance on live microscopy and spatial-omics data
Miao, Yang et al.
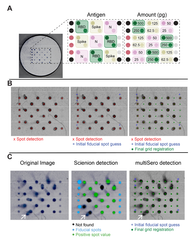
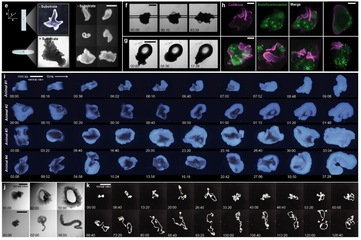
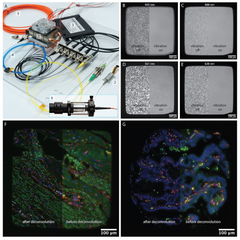
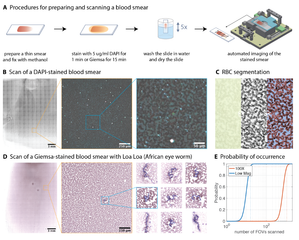
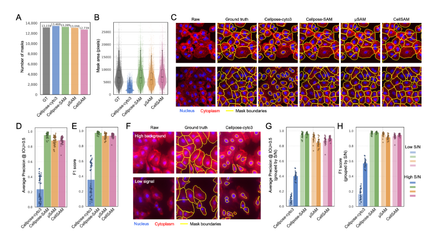
Systematic benchmark of generalist deep-learning segmentation models—Cellpose cyto3, Cellpose-SAM, μSAM, CellSAM, Mesmer, and InstanSeg—across phase contrast, fluorescence cell culture, and multiplexed CODEX tissue imaging. Cellpose-SAM excels on phase contrast and SAM-based models on fluorescence, while no single model dominates on CODEX, with downstream clustering and cell-type identification varying by choice. The work argues for selecting segmentation models based on dataset characteristics and analytical goals rather than a one-size-fits-all approach.